 We provide a concise workflow for the selection of the best assembly tool. We found that assembler choice ultimately depends on the scientific question, the available resources and the bioinformatic competence of the researcher. MEGAHIT emerged as a computationally inexpensive alternative to SPAdes, assembling the most complex dataset using less than 500 GB of RAM and within 10 hours. Overall, we found that SPAdes provided the largest contigs and highest N50 values across 6 of the 9 environmental datasets, followed by MEGAHIT and metaSPAdes. To assist with selection of an appropriate metagenome assembler we evaluated the capabilities of nine prominent assembly tools on nine publicly-available environmental metagenomes, as well as three simulated datasets. However, while several platforms have been developed for this critical step, there is currently no clear framework for the assembly of metagenomic sequence data. The analysis of metagenomic sequences facilitates gene prediction and annotation, and enables the assembly of draft genomes, including uncultured members of a community. Supplementary_files_format_and_content: Tab-delimited table format containing gene names (annotated by HGNC-approved gene symbol) and nomalized RPKM values for all samples.Īnalysis of the expression profiles of back skin and excisional wound-derived fibroblasts FACS-sorted from mice overexpressing activin A.Metagenomics allows unprecedented access to uncultured environmental microorganisms. Differential expression analysis was performed across all group comparisons using the edgeR exact test as implemented in the CLC Genomics Workbench (Qiagen), and differentially expressed genes (DEGs) were determined and compared. Principal component analysis (PCA) was performed to evaluate grouping of samples. Numbers of RNA transcripts were determined on the gene level and normalized to reads per kilobase of transcript, per million mapped reads (RPKM). Quality control via CLC Genomics Workbench excluded one sample (rep2) from the Act_5DW group all other samples generated 20-40 million reads, showed high (82.3-95.7%) mapping rates to the mouse reference genome, high (93.7-97.5%) mapping rates to genes and low (0.2-6.1%) rRNA mapping rates. 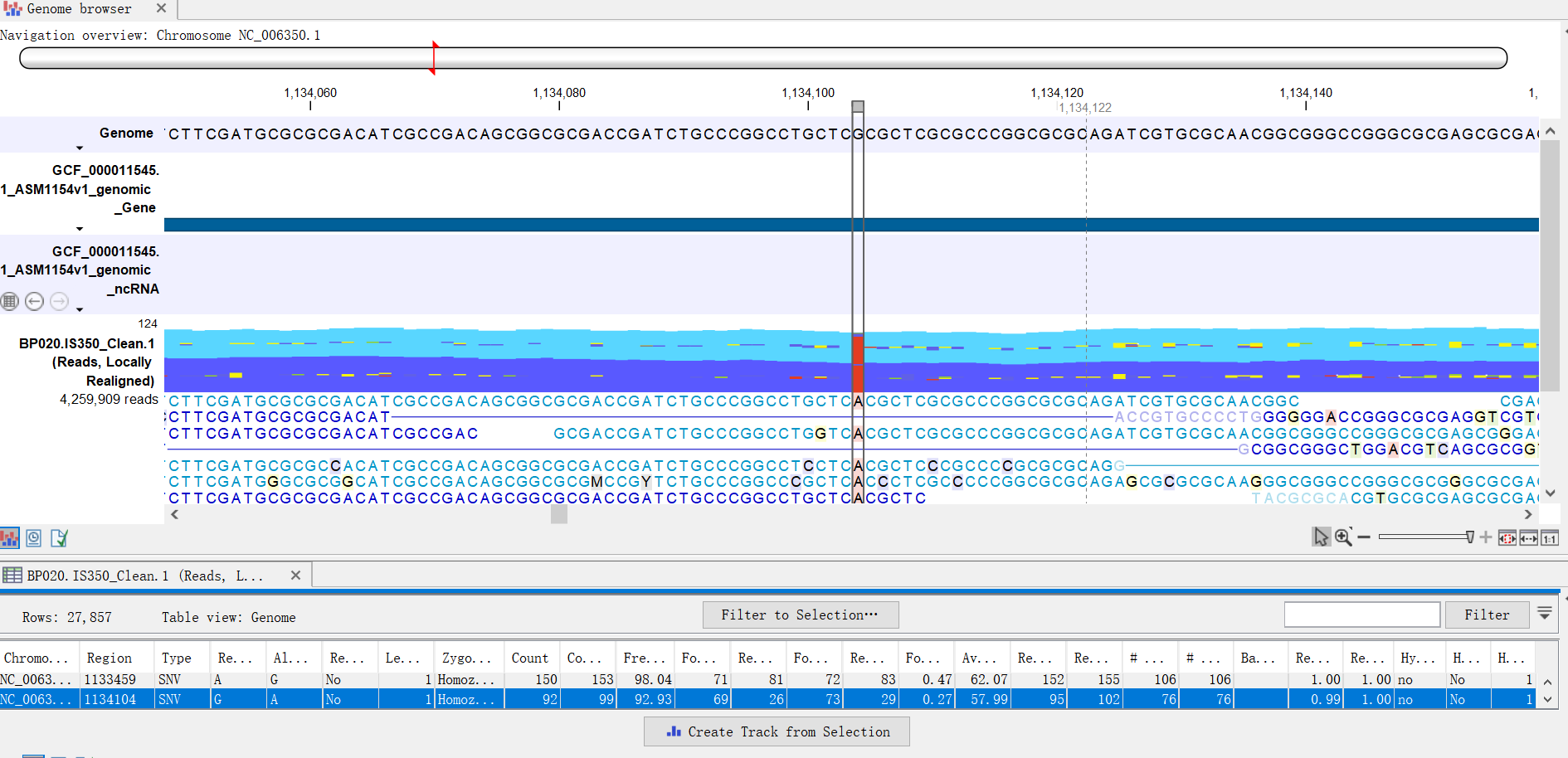
Sequence alignment to the Mus musculus reference genome (build GRCm38) and transcript expression quantification of the resulting high-quality reads was performed using the CLC Genomics Workbench (Qiagen). RNA was analyzed via TapeStation (Agilent), and samples with RIN>6.8 were adjusted to 100 ng (back skin) or 200 ng (5-day wounds).Įxtracted RNA samples were subjected to the RNA sequencing protocol via poly-A enrichment, True-Seq library preparation, and stranded 100bp sequencing on an Illumina HiSeq 2500 instrument. Total RNA was isolated using the RNeasy Micro Kit (Qiagen). Cells were sorted into cold empty Eppendorf tubes with FACS buffer containing RNAase inhibitor and centrifuged for total RNA extraction. Dead cells were excluded by Sytox Green staining. CD3, CD11b and F4/80 were used as additional negative markers. Back skin and 5-day excisional wound fibroblasts were enriched by FACS sorting of CD45-negative/CD140a-positive (CD45-CD140a+) cells.  
Following digestion, the cell suspension was filtered through a 70 µm nylon strainer, centrifuged at 300g at 4☌ for 10 min, and resuspended in FACS buffer. GEO help: Mouse over screen elements for information.Ĭell type/source: FACS-sorted CD45-CD140a+ live cellsīack skin and 5-day wounds were harvested, minced with scissors, and digested with 0.25 mg/ml Liberase TL (Roche) in DMEM for 1 h at 37☌. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
